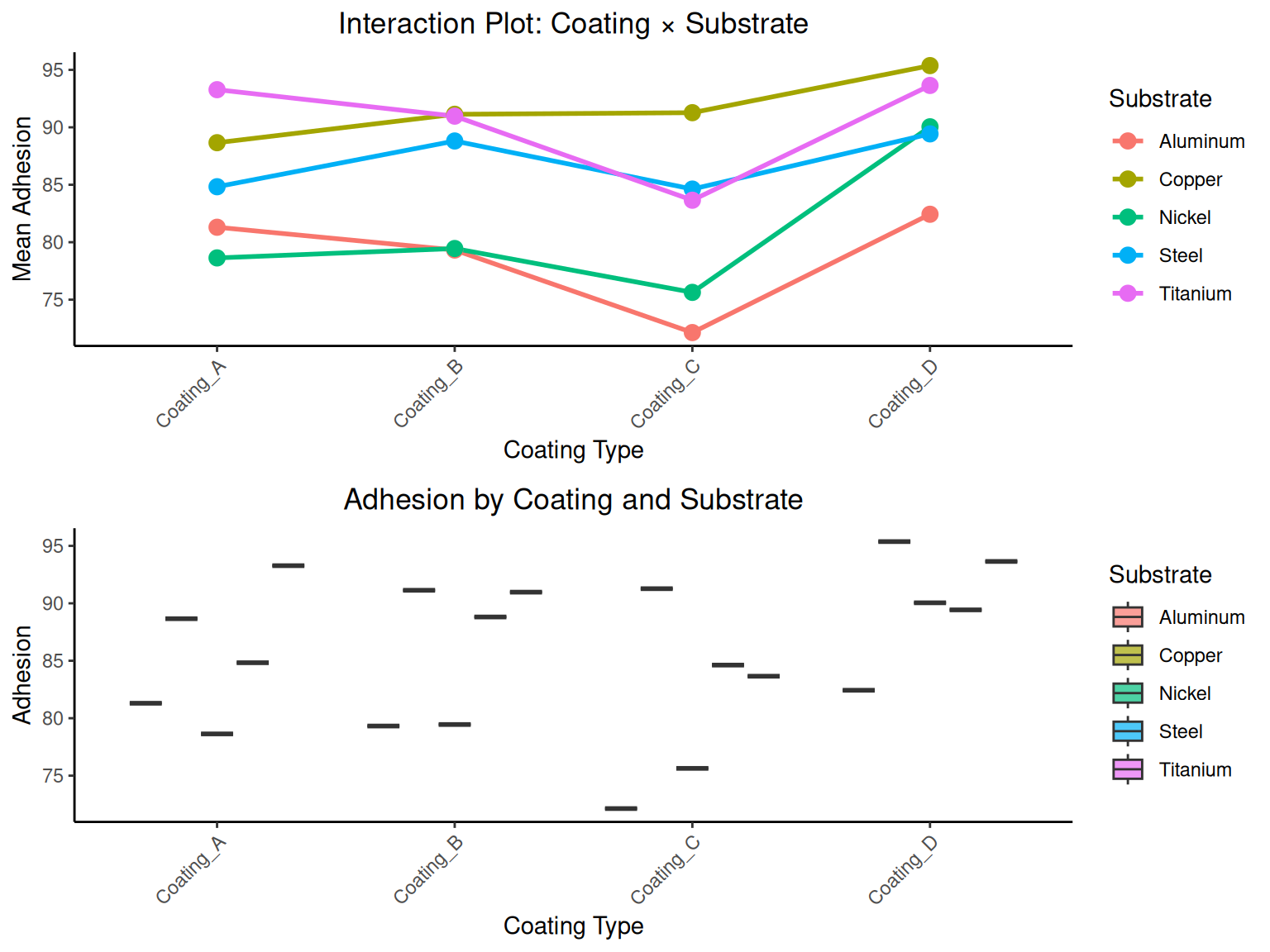
# RCBD ANOVA calculations
a_rcbd <- length(unique(coating_data$coating)) # treatments
b_rcbd <- length(unique(coating_data$substrate)) # blocks
N_rcbd <- fnobs(coating_data$adhesion)
# Calculate sum of squares for RCBD
grand_mean_rcbd <- fmean(coating_data$adhesion)
# Treatment sum of squares
treatment_means <- coating_summary$Mean
SS_treatments_rcbd <- b_rcbd * sum((treatment_means - grand_mean_rcbd)^2)
# Block sum of squares
block_means <- substrate_summary$Mean
SS_blocks_rcbd <- a_rcbd * sum((block_means - grand_mean_rcbd)^2)
# Total sum of squares
SS_total_rcbd <- sum((coating_data$adhesion - grand_mean_rcbd)^2)
# Error sum of squares (by subtraction)
SS_error_rcbd <- SS_total_rcbd - SS_treatments_rcbd - SS_blocks_rcbd
# Degrees of freedom
df_treatments_rcbd <- a_rcbd - 1
df_blocks_rcbd <- b_rcbd - 1
df_error_rcbd <- (a_rcbd - 1) * (b_rcbd - 1)
df_total_rcbd <- N_rcbd - 1
# Mean squares
MS_treatments_rcbd <- SS_treatments_rcbd / df_treatments_rcbd
MS_blocks_rcbd <- SS_blocks_rcbd / df_blocks_rcbd
MS_error_rcbd <- SS_error_rcbd / df_error_rcbd
# F-statistics
f_treatments_rcbd <- MS_treatments_rcbd / MS_error_rcbd
f_blocks_rcbd <- MS_blocks_rcbd / MS_error_rcbd
# P-values
p_treatments_rcbd <- 1 - pf(f_treatments_rcbd, df_treatments_rcbd, df_error_rcbd)
p_blocks_rcbd <- 1 - pf(f_blocks_rcbd, df_blocks_rcbd, df_error_rcbd)
# RCBD ANOVA table
rcbd_anova_table <- data.table(
Source = c("Coatings", "Substrates", "Error", "Total"),
SS = c(round(SS_treatments_rcbd, 2), round(SS_blocks_rcbd, 2),
round(SS_error_rcbd, 2), round(SS_total_rcbd, 2)),
df = c(df_treatments_rcbd, df_blocks_rcbd, df_error_rcbd, df_total_rcbd),
MS = c(round(MS_treatments_rcbd, 2), round(MS_blocks_rcbd, 2),
round(MS_error_rcbd, 2), NA),
F_Statistic = c(round(f_treatments_rcbd, 4), round(f_blocks_rcbd, 4), NA, NA),
P_Value = c(round(p_treatments_rcbd, 4), round(p_blocks_rcbd, 4), NA, NA)
)
print("RCBD ANOVA Table:")
print(rcbd_anova_table)
# RCBD conclusions
rcbd_conclusions <- data.table(
Effect = c("Coating Effect", "Substrate Effect"),
F_Statistic = c(round(f_treatments_rcbd, 4), round(f_blocks_rcbd, 4)),
P_Value = c(round(p_treatments_rcbd, 4), round(p_blocks_rcbd, 4)),
Decision = c(
ifelse(p_treatments_rcbd < alpha_anova, "Reject H0", "Fail to reject H0"),
ifelse(p_blocks_rcbd < alpha_anova, "Reject H0", "Fail to reject H0")
),
Conclusion = c(
ifelse(p_treatments_rcbd < alpha_anova, "Coating means differ significantly", "No significant coating effect"),
ifelse(p_blocks_rcbd < alpha_anova, "Substrate means differ significantly", "No significant substrate effect")
)
)
print("RCBD Test Results:")
print(rcbd_conclusions)
# Using R's aov for verification
rcbd_r <- aov(adhesion ~ coating + substrate, data = coating_data)
rcbd_summary_r <- summary(rcbd_r)
print("R's RCBD ANOVA verification:")
print(rcbd_summary_r)