An introduction to R for Agricultural Research
- Introduction
- Regression Analysis
- Simple Linear Regression
- Multiple Linear Regression
- Polynomial Regression Analysis
- Analysis of Variance (ANOVA)
- Analysis of Covariance (ANCOVA)
- Correlation Analysis
- Simple Correlation Analysis
- Partial Correlation Analysis
- Multiple Correlation Analysis
- Completely Randomized Design (CRD)
- Randomized Complete Block Design (RCBD)
- Latin Square Design
Introduction
R is a free, open-source programming language and software environment for statistical computing, bioinformatics, visualization and general computing. R provides a wide variety of statistical and graphical techniques, and is highly extensible. The latest version of R is 3.5.0 which can be obtained from https://cran.r-project.org/bin/windows/base/.
R R-3.5.0 for Windows (32/64 bit)
RStudio RStudio
Manual An Introduction to R
Ref Card R Reference Card
New York Times New York Times
Nature Article Nature Article
Regression Analysis
- Quantifying dependency of a normal response on quantitative explanatory variable(s)
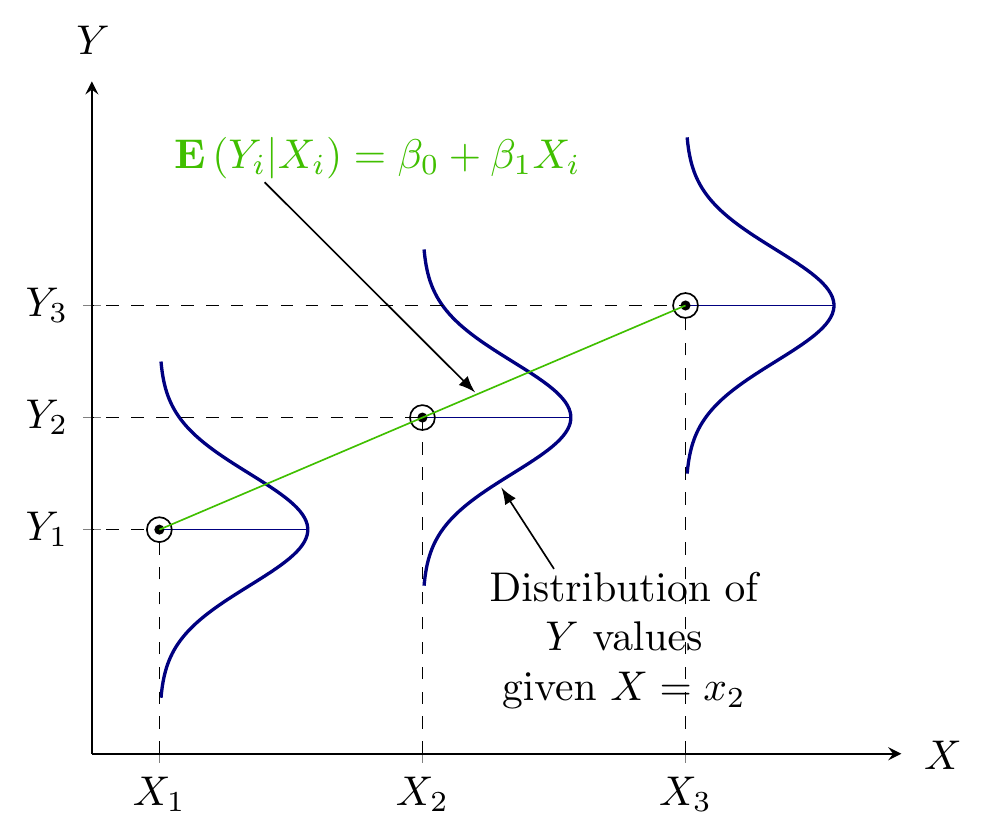
Figure 1: Population Regression Function
Simple Linear Regression
- Quantifying dependency of a normal response on a quantitative explanatory variable
Example
| Fertilizer (Kg/acre) | Production (Bushels/acre) |
|---|---|
| 100 | 70 |
| 200 | 70 |
| 400 | 80 |
| 500 | 100 |
# Fertilizer (Kg/acre)
# Production (Bushels/acre)
Fertilizer <- c(100, 200, 400, 500)
Production <- c( 70, 70, 80, 100)
df1 <- data.frame(Fertilizer, Production)
df1 Fertilizer Production
1 100 70
2 200 70
3 400 80
4 500 100library(ggplot2)
# if (!require("ggplot2")) install.packages("ggplot2")
p1 <-
ggplot(data = df1, mapping = aes(x = Fertilizer, y = Production)) +
geom_point() +
labs(x = "Fertilizer", y = "Production") +
theme_bw()
print(p1)fm1 <- lm(formula = Production ~ Fertilizer, data = df1)
summary(fm1)
Call:
lm(formula = Production ~ Fertilizer, data = df1)
Residuals:
1 2 3 4
4 -3 -7 6
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 59.00000 7.95299 7.419 0.0177 *
Fertilizer 0.07000 0.02345 2.985 0.0963 .
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 7.416 on 2 degrees of freedom
Multiple R-squared: 0.8167, Adjusted R-squared: 0.725
F-statistic: 8.909 on 1 and 2 DF, p-value: 0.0963anova(fm1)Analysis of Variance Table
Response: Production
Df Sum Sq Mean Sq F value Pr(>F)
Fertilizer 1 490 490 8.9091 0.0963 .
Residuals 2 110 55
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Example
| Fertilizer | Yield |
|---|---|
| 0.3 | 10 |
| 0.6 | 15 |
| 0.9 | 30 |
| 1.2 | 35 |
| 1.5 | 25 |
| 1.8 | 30 |
| 2.1 | 50 |
| 2.4 | 45 |
Fert <- c(0.3, 0.6, 0.9, 1.2, 1.5, 1.8, 2.1, 2.4)
Yield <- c(10, 15, 30, 35, 25, 30, 50, 45)
df2 <- data.frame(Fert, Yield)
df2 Fert Yield
1 0.3 10
2 0.6 15
3 0.9 30
4 1.2 35
5 1.5 25
6 1.8 30
7 2.1 50
8 2.4 45library(ggplot2)
# if (!require("ggplot2")) install.packages("ggplot2")
p2 <-
ggplot(data = df2, mapping = aes(x = Fert, y = Yield)) +
geom_point() +
labs(x = "Fertilizer", y = "Yield") +
theme_bw()
print(p2)fm2 <- lm(formula = Yield ~ Fert, data = df2)
summary(fm2)
Call:
lm(formula = Yield ~ Fert, data = df2)
Residuals:
Min 1Q Median 3Q Max
-7.441 -4.018 -2.441 7.351 7.798
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 8.036 5.504 1.460 0.1946
Fert 16.270 3.633 4.478 0.0042 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 7.064 on 6 degrees of freedom
Multiple R-squared: 0.7697, Adjusted R-squared: 0.7313
F-statistic: 20.05 on 1 and 6 DF, p-value: 0.004202anova(fm2)Analysis of Variance Table
Response: Yield
Df Sum Sq Mean Sq F value Pr(>F)
Fert 1 1000.6 1000.6 20.052 0.004202 **
Residuals 6 299.4 49.9
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Example
| Weekly Income ($) | Weekly Expenditures ($) |
|---|---|
| 80 | 70 |
| 100 | 65 |
| 120 | 90 |
| 140 | 95 |
| 160 | 110 |
| 180 | 115 |
| 200 | 120 |
| 220 | 140 |
| 240 | 155 |
| 260 | 150 |
# Weekly Income ($) of a Family
# Weekly Expenditures ($) of a Family
Income <- seq(from = 80, to = 260, by = 20)
Expenditures <- c(70, 65, 90, 95, 110, 115, 120, 140, 155, 150)
df3 <- data.frame(Income, Expenditures)
df3 Income Expenditures
1 80 70
2 100 65
3 120 90
4 140 95
5 160 110
6 180 115
7 200 120
8 220 140
9 240 155
10 260 150library(ggplot2)
# if (!require("ggplot2")) install.packages("ggplot2")
p3 <-
ggplot(data = df3, mapping = aes(x = Income, y = Expenditures)) +
geom_point() +
scale_x_continuous(labels = scales::dollar) +
scale_y_continuous(labels = scales::dollar) +
labs(x = "Weekly income, $", y = "Weekly consumption expenditures, $") +
theme_bw()
print(p3)fm3 <- lm(formula = Expenditures ~ Income, data = df3)
summary(fm3)
Call:
lm(formula = Expenditures ~ Income, data = df3)
Residuals:
Min 1Q Median 3Q Max
-10.364 -4.977 1.409 4.364 8.364
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 24.45455 6.41382 3.813 0.00514 **
Income 0.50909 0.03574 14.243 5.75e-07 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 6.493 on 8 degrees of freedom
Multiple R-squared: 0.9621, Adjusted R-squared: 0.9573
F-statistic: 202.9 on 1 and 8 DF, p-value: 5.753e-07anova(fm3)Analysis of Variance Table
Response: Expenditures
Df Sum Sq Mean Sq F value Pr(>F)
Income 1 8552.7 8552.7 202.87 5.753e-07 ***
Residuals 8 337.3 42.2
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Multiple Linear Regression
- Quantifying dependency of a normal response on two or more quantitative explanatory variables
Example
| Fertilizer (Kg) | Rainfall (mm) | Yield (Kg) |
|---|---|---|
| 100 | 10 | 40 |
| 200 | 20 | 50 |
| 300 | 10 | 50 |
| 400 | 30 | 70 |
| 500 | 20 | 65 |
| 600 | 20 | 65 |
| 700 | 30 | 80 |
# Fertilizer (Kg)
# Rainfall (mm)
# Yield (Kg)
Fertilizer <- seq(from = 100, to = 700, by = 100)
Rainfall <- c(10, 20, 10, 30, 20, 20, 30)
Yield <- c(40, 50, 50, 70, 65, 65, 80)
df4 <- data.frame(Fertilizer, Rainfall, Yield)
df4 Fertilizer Rainfall Yield
1 100 10 40
2 200 20 50
3 300 10 50
4 400 30 70
5 500 20 65
6 600 20 65
7 700 30 80library(car)
# if (!require("car")) install.packages("car")
car::scatter3d(formula = Yield ~ Fertilizer + Rainfall, data = df4)fm4 <- lm(formula = Yield ~ Fertilizer + Rainfall, data = df4)
summary(fm4)
Call:
lm(formula = Yield ~ Fertilizer + Rainfall, data = df4)
Residuals:
1 2 3 4 5 6 7
-0.2381 -2.3810 2.1429 1.6667 1.1905 -2.6190 0.2381
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 28.095238 2.491482 11.277 0.000352 ***
Fertilizer 0.038095 0.005832 6.532 0.002838 **
Rainfall 0.833333 0.154303 5.401 0.005690 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 2.315 on 4 degrees of freedom
Multiple R-squared: 0.9814, Adjusted R-squared: 0.972
F-statistic: 105.3 on 2 and 4 DF, p-value: 0.0003472anova(fm4)Analysis of Variance Table
Response: Yield
Df Sum Sq Mean Sq F value Pr(>F)
Fertilizer 1 972.32 972.32 181.500 0.0001756 ***
Rainfall 1 156.25 156.25 29.167 0.0056899 **
Residuals 4 21.43 5.36
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1library(lm.beta)
# if (!require("lm.beta")) install.packages("lm.beta")
lm.beta(fm4)
Call:
lm(formula = Yield ~ Fertilizer + Rainfall, data = df4)
Standardized Coefficients::
(Intercept) Fertilizer Rainfall
0.0000000 0.5944301 0.4914732
Polynomial Regression Analysis
- Quantifying non-linear dependency of a normal response on quantitative explanatory variable(s)
Example
An experiment was conducted to evaluate the effects of different levels of nitrogen. Three levels of nitrogen: 0, 10 and 20 grams per plot were used in the experiment. Each treatment was replicated twice and data is given below:
| Nitrogen | Yield |
|---|---|
| 0 | 5 |
| 0 | 7 |
| 10 | 15 |
| 10 | 17 |
| 20 | 9 |
| 20 | 11 |
Nitrogen <- c(0, 0, 10, 10, 20, 20)
Yield <- c(5, 7, 15, 17, 9, 11)
df5 <- data.frame(Nitrogen, Yield)
df5 Nitrogen Yield
1 0 5
2 0 7
3 10 15
4 10 17
5 20 9
6 20 11library(ggplot2)
# if (!require("ggplot2")) install.packages("ggplot2")
p5 <-
ggplot(data = df5, mapping = aes(x = Nitrogen, y = Yield)) +
geom_point() +
labs(x = "Nitrogen", y = "Yield") +
theme_bw()
print(p5)fm5 <- lm(formula = Yield ~ Nitrogen + I(Nitrogen^2), data = df5)
# fm5 <- lm(formula = Yield ~ poly(x = Nitrogen, degree = 2, raw = TRUE), data = df5)
summary(fm5)
Call:
lm(formula = Yield ~ Nitrogen + I(Nitrogen^2), data = df5)
Residuals:
1 2 3 4 5 6
-1 1 -1 1 -1 1
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 6.00000 1.00000 6.000 0.00927 **
Nitrogen 1.80000 0.25495 7.060 0.00584 **
I(Nitrogen^2) -0.08000 0.01225 -6.532 0.00729 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 1.414 on 3 degrees of freedom
Multiple R-squared: 0.9441, Adjusted R-squared: 0.9068
F-statistic: 25.33 on 2 and 3 DF, p-value: 0.01322anova(fm5)Analysis of Variance Table
Response: Yield
Df Sum Sq Mean Sq F value Pr(>F)
Nitrogen 1 16.000 16.000 8.000 0.066276 .
I(Nitrogen^2) 1 85.333 85.333 42.667 0.007292 **
Residuals 3 6.000 2.000
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Analysis of Variance (ANOVA)
- Comparing means of Normal dependent variable for levels of different factor(s)
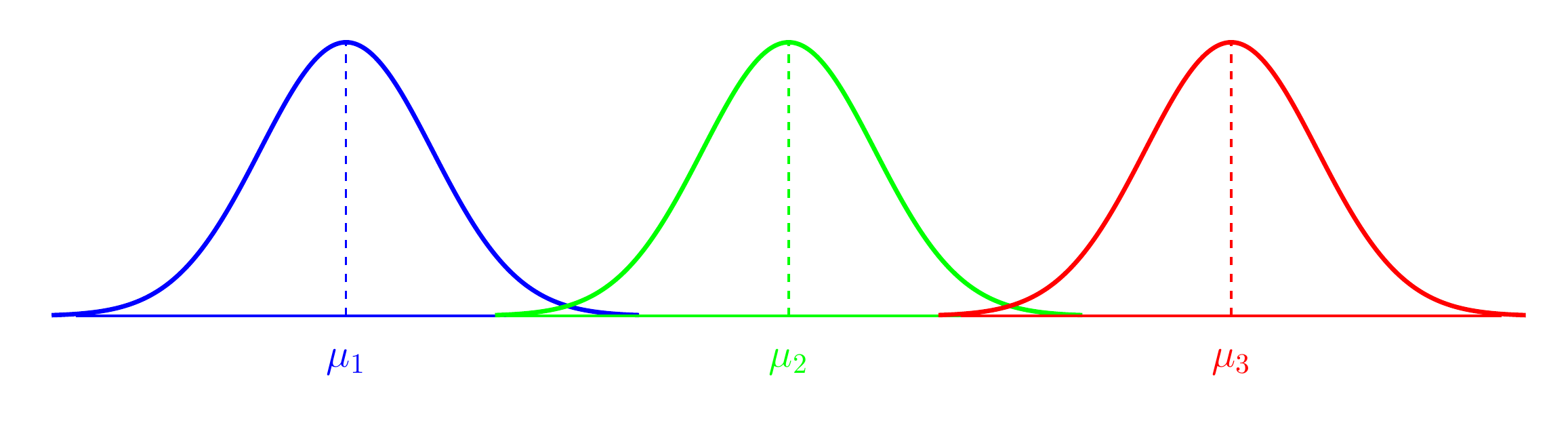
Figure 2: Analysis of Variance
Example
| Yield | Variety |
|---|---|
| 5 | V1 |
| 6 | V1 |
| 7 | V1 |
| 15 | V2 |
| 16 | V2 |
| 17 | V2 |
Yield <- c(5, 6, 7, 15, 16, 17)
Variety <- gl(n = 2, k = 3, length = 2*3, labels = c("V1", "V2"))
df6 <- data.frame(Yield, Variety)
df6 Yield Variety
1 5 V1
2 6 V1
3 7 V1
4 15 V2
5 16 V2
6 17 V2library(ggplot2)
# if (!require("ggplot2")) install.packages("ggplot2")
p6 <-
ggplot(data = df6, mapping = aes(x = Variety, y = Yield)) +
geom_point() +
geom_boxplot() +
labs(x = "Variety", y = "Yield") +
theme_bw()
print(p6)fm6 <- lm(formula = Yield ~ Variety, data = df6)
summary(fm6)
Call:
lm(formula = Yield ~ Variety, data = df6)
Residuals:
1 2 3 4 5 6
-1.000e+00 -4.774e-15 1.000e+00 -1.000e+00 -1.110e-16 1.000e+00
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 6.0000 0.5774 10.39 0.000484 ***
VarietyV2 10.0000 0.8165 12.25 0.000255 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 1 on 4 degrees of freedom
Multiple R-squared: 0.974, Adjusted R-squared: 0.9675
F-statistic: 150 on 1 and 4 DF, p-value: 0.0002552anova(fm6)Analysis of Variance Table
Response: Yield
Df Sum Sq Mean Sq F value Pr(>F)
Variety 1 150 150 150 0.0002552 ***
Residuals 4 4 1
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Example
| Yield | Variety |
|---|---|
| 5 | V1 |
| 7 | V1 |
| 15 | V2 |
| 17 | V2 |
| 17 | V3 |
| 19 | V3 |
Yield <- c(5, 7, 15, 17, 17, 19)
Variety <- gl(n = 3, k = 2, length = 3*2, labels = c("V1", "V2", "V3"))
df7 <- data.frame(Yield, Variety)
df7 Yield Variety
1 5 V1
2 7 V1
3 15 V2
4 17 V2
5 17 V3
6 19 V3library(ggplot2)
# if (!require("ggplot2")) install.packages("ggplot2")
p7 <-
ggplot(data = df7, mapping = aes(x = Variety, y = Yield)) +
geom_point() +
geom_boxplot() +
labs(x = "Variety", y = "Yield") +
theme_bw()
print(p7)fm7 <- lm(formula = Yield ~ Variety, data = df7)
summary(fm7)
Call:
lm(formula = Yield ~ Variety, data = df7)
Residuals:
1 2 3 4 5 6
-1 1 -1 1 -1 1
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 6.000 1.000 6.000 0.00927 **
VarietyV2 10.000 1.414 7.071 0.00582 **
VarietyV3 12.000 1.414 8.485 0.00344 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 1.414 on 3 degrees of freedom
Multiple R-squared: 0.965, Adjusted R-squared: 0.9416
F-statistic: 41.33 on 2 and 3 DF, p-value: 0.006553anova(fm7)Analysis of Variance Table
Response: Yield
Df Sum Sq Mean Sq F value Pr(>F)
Variety 2 165.33 82.667 41.333 0.006553 **
Residuals 3 6.00 2.000
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Analysis of Covariance (ANCOVA)
- Quantifying dependency of a normal response on quantitative explanatory variable(s)
- Comparing means of Normal dependent variable for levels of different factor(s)
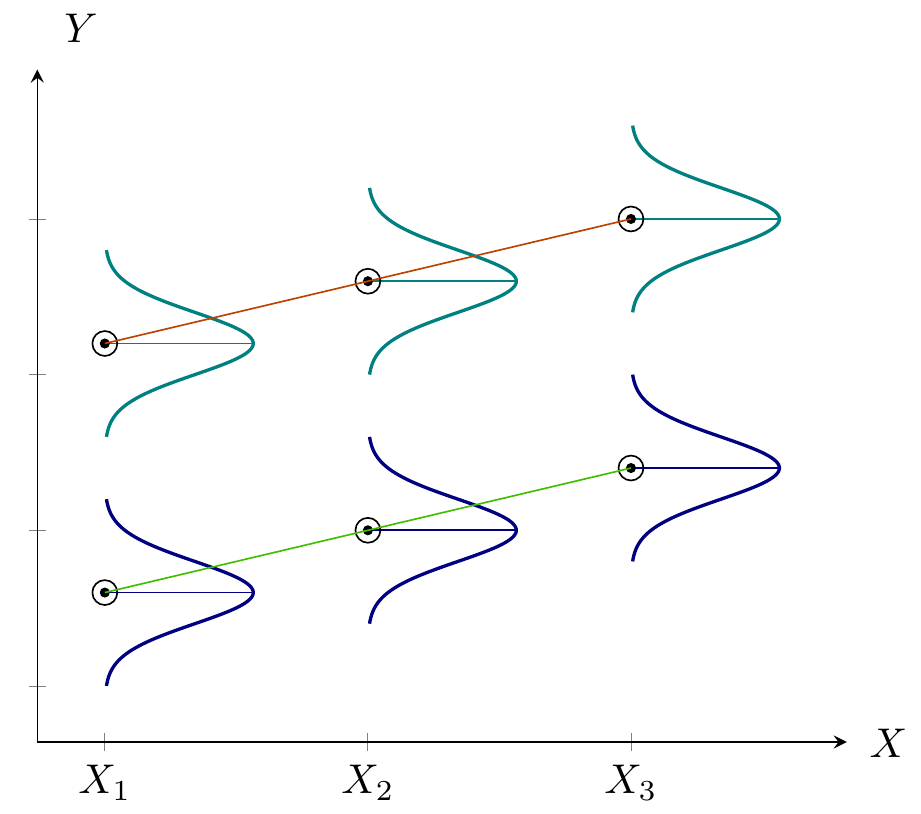
Figure 3: Analysis of Covariance
Example
| Yield | Fert | Variety |
|---|---|---|
| 51 | 80 | V1 |
| 52 | 80 | V1 |
| 53 | 90 | V1 |
| 54 | 90 | V1 |
| 56 | 100 | V1 |
| 57 | 100 | V1 |
| 55 | 80 | V2 |
| 56 | 80 | V2 |
| 58 | 90 | V2 |
| 59 | 90 | V2 |
| 62 | 100 | V2 |
| 63 | 100 | V2 |
Yield <- c(51, 52, 53, 54, 56, 57, 55, 56, 58, 59, 62, 63)
Fert <- rep(x = c(80, 90, 100), each = 2)
Variety <- gl(n = 2, k = 6, length = 2*6, labels = c("V1", "V2"), ordered = FALSE)
df8 <- data.frame(Yield, Fert, Variety)
df8 Yield Fert Variety
1 51 80 V1
2 52 80 V1
3 53 90 V1
4 54 90 V1
5 56 100 V1
6 57 100 V1
7 55 80 V2
8 56 80 V2
9 58 90 V2
10 59 90 V2
11 62 100 V2
12 63 100 V2Same intercepts but different slopes
fm8 <- lm(formula = Yield ~ Fert + Variety, data = df8)
summary(fm8)
Call:
lm(formula = Yield ~ Fert + Variety, data = df8)
Residuals:
Min 1Q Median 3Q Max
-0.8333 -0.8333 0.1667 0.1667 1.1667
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 26.83333 2.54557 10.54 2.30e-06 ***
Fert 0.30000 0.02805 10.69 2.04e-06 ***
VarietyV2 5.00000 0.45812 10.91 1.72e-06 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.7935 on 9 degrees of freedom
Multiple R-squared: 0.9629, Adjusted R-squared: 0.9546
F-statistic: 116.7 on 2 and 9 DF, p-value: 3.657e-07anova(fm8)Analysis of Variance Table
Response: Yield
Df Sum Sq Mean Sq F value Pr(>F)
Fert 1 72.000 72.00 114.35 2.042e-06 ***
Variety 1 75.000 75.00 119.12 1.720e-06 ***
Residuals 9 5.667 0.63
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1library(ggplot2)
# if (!require("ggplot2")) install.packages("ggplot2")
p8 <-
ggplot(data = df8, mapping = aes(x = Fert, y = Yield, color = Variety)) +
geom_point() +
geom_line(mapping = aes(y = predict(fm8))) +
labs(x = "Fert", y = "Yield") +
theme_bw()
print(p8)
Different intercepts and different slopes
fm9 <- lm(formula = Yield ~ Fert * Variety, data = df8)
summary(fm9)
Call:
lm(formula = Yield ~ Fert * Variety, data = df8)
Residuals:
Min 1Q Median 3Q Max
-0.83333 -0.33333 -0.08333 0.66667 0.66667
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 31.33333 3.05903 10.243 7.09e-06 ***
Fert 0.25000 0.03385 7.385 7.73e-05 ***
VarietyV2 -4.00000 4.32612 -0.925 0.3822
Fert:VarietyV2 0.10000 0.04787 2.089 0.0701 .
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 0.677 on 8 degrees of freedom
Multiple R-squared: 0.976, Adjusted R-squared: 0.967
F-statistic: 108.4 on 3 and 8 DF, p-value: 8.11e-07anova(fm9)Analysis of Variance Table
Response: Yield
Df Sum Sq Mean Sq F value Pr(>F)
Fert 1 72.000 72.000 157.0909 1.538e-06 ***
Variety 1 75.000 75.000 163.6364 1.315e-06 ***
Fert:Variety 1 2.000 2.000 4.3636 0.07013 .
Residuals 8 3.667 0.458
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1library(ggplot2)
# if (!require("ggplot2")) install.packages("ggplot2")
p9 <-
ggplot(data = df8, mapping = aes(x = Fert, y = Yield, color = Variety)) +
geom_point() +
geom_line(mapping = aes(y = predict(fm9))) +
labs(x = "Fert", y = "Yield") +
theme_bw()
print(p9)Correlation Analysis
- Linear Relationship between Quantitative Variables
Simple Correlation Analysis
- Linear Relationship between Two Quantitative Variables
- \(\left(X_{1},X_{2}\right)\)
Example
| Sparrow Wing length (cm) | Sparrow Tail length (cm) |
|---|---|
| 10.4 | 7.4 |
| 10.8 | 7.6 |
| 11.1 | 7.9 |
| 10.2 | 7.2 |
| 10.3 | 7.4 |
| 10.2 | 7.1 |
| 10.7 | 7.4 |
| 10.5 | 7.2 |
| 10.8 | 7.8 |
| 11.2 | 7.7 |
| 10.6 | 7.8 |
| 11.4 | 8.3 |
WingLength <- c(10.4, 10.8, 11.1, 10.2, 10.3, 10.2, 10.7, 10.5, 10.8, 11.2, 10.6, 11.4)
TailLength <- c(7.4, 7.6, 7.9, 7.2, 7.4, 7.1, 7.4, 7.2, 7.8, 7.7, 7.8, 8.3)
df10 <- data.frame(WingLength, TailLength)
df10 WingLength TailLength
1 10.4 7.4
2 10.8 7.6
3 11.1 7.9
4 10.2 7.2
5 10.3 7.4
6 10.2 7.1
7 10.7 7.4
8 10.5 7.2
9 10.8 7.8
10 11.2 7.7
11 10.6 7.8
12 11.4 8.3library(ggplot2)
# if (!require("ggplot2")) install.packages("ggplot2")
p10 <-
ggplot(data = df10, mapping = aes(x= WingLength, y = TailLength)) +
geom_point() +
labs(x = "Sparrow Wing Length (cm)", y = "Sparrow Tail Length (cm)") +
theme_bw()
print(p10)cor(df10$WingLength, df10$TailLength)[1] 0.8703546cor.test(df10$WingLength, df10$TailLength)
Pearson's product-moment correlation
data: df10$WingLength and df10$TailLength
t = 5.5893, df = 10, p-value = 0.0002311
alternative hypothesis: true correlation is not equal to 0
95 percent confidence interval:
0.5923111 0.9631599
sample estimates:
cor
0.8703546
Partial Correlation Analysis
- Linear Relationship between Quantitative Variables while holding/keeping all other constants
- \(\left(X_{1},X_{2}\right)|X_{3}\)
Example
| Leaf Area (cm^2) | Leaf Moisture (%) | Total Shoot Length (cm) |
|---|---|---|
| 72 | 80 | 307 |
| 174 | 75 | 529 |
| 116 | 81 | 632 |
| 78 | 83 | 527 |
| 134 | 79 | 442 |
| 95 | 81 | 525 |
| 113 | 80 | 481 |
| 98 | 81 | 710 |
| 148 | 74 | 422 |
| 42 | 78 | 345 |
LeafArea <- c(72, 174, 116, 78, 134, 95, 113, 98, 148, 42)
LeafMoist <- c(75, 81, 83, 79, 81, 80, 81, 74, 78, 58)
TotShLength <- c(307, 529, 632, 527, 442, 525, 481, 710, 422, 345)
df11 <- data.frame(LeafArea, LeafMoist, TotShLength)
df11 LeafArea LeafMoist TotShLength
1 72 75 307
2 174 81 529
3 116 83 632
4 78 79 527
5 134 81 442
6 95 80 525
7 113 81 481
8 98 74 710
9 148 78 422
10 42 58 345library(ppcor)
# if (!require("ppcor")) install.packages("ppcor")
pcor(df11)$estimate
LeafArea LeafMoist TotShLength
LeafArea 1.000000000 0.6509096 -0.007012236
LeafMoist 0.650909618 1.0000000 0.324334659
TotShLength -0.007012236 0.3243347 1.000000000
$p.value
LeafArea LeafMoist TotShLength
LeafArea 0.00000000 0.05760462 0.9857154
LeafMoist 0.05760462 0.00000000 0.3944839
TotShLength 0.98571537 0.39448385 0.0000000
$statistic
LeafArea LeafMoist TotShLength
LeafArea 0.00000000 2.268502 -0.01855309
LeafMoist 2.26850175 0.000000 0.90714704
TotShLength -0.01855309 0.907147 0.00000000
$n
[1] 10
$gp
[1] 1
$method
[1] "pearson"
Multiple Correlation Analysis
- Linear Relationship between a Quantitative Variable and set of other Quantitative Variables
- \(\left(X_{1},\left[X_{2},X_{3}\right]\right)\)
Example
| Leaf Area (cm^2) | Leaf Moisture (%) | Total Shoot Length (cm) |
|---|---|---|
| 72 | 80 | 307 |
| 174 | 75 | 529 |
| 116 | 81 | 632 |
| 78 | 83 | 527 |
| 134 | 79 | 442 |
| 95 | 81 | 525 |
| 113 | 80 | 481 |
| 98 | 81 | 710 |
| 148 | 74 | 422 |
| 42 | 78 | 345 |
fm12 <- lm(formula = TotShLength ~ LeafArea + LeafMoist, data = df11)
sqrt(summary(fm12)$r.squared)[1] 0.421277
Completely Randomized Design (CRD)
- Used when experimental material is homogeneous
Example
The following table shows some of the results of an experiment on the effects of applications of sulphur in reducing scab disease of potatoes. The object in applying sulphur is to increase the acidity of the soil, since scab does not thrive in very acid soil. In addition to untreated plots which serve as a control, 3 amounts of dressing were compared—300, 600, and 900 lb. per acre. Both a fall and a spring application of each amount was tested, so that in all there were seven distinct treatments. The sulphur was spread by hand on the surface of the soil, and then diced into a depth of about 4 inches. The quantity to be analyzed is the “scab index”. That is roughly speaking, the percentage of the surface area of the potato that is infected with scab. It is obtained by examining 100 potatoes at random from each plot, grading each potato on a scale from 0 to 100% infected, and taking the average.
ScabIndex <-
c(
12, 10, 24, 29,
30, 18, 32, 26,
9, 9, 16, 4,
30, 7, 21, 9,
16, 10, 18, 18,
18, 24, 12, 19,
10, 4, 4, 5,
17, 7, 16, 17
)
Trt <- factor(
x = rep(x = 1:7, times = c(8, 4, 4, 4, 4, 4, 4))
, labels = c("O", "F3", "S3", "F6", "S6", "F9", "S9")
)
df13 <- data.frame(Trt, ScabIndex)
df13 Trt ScabIndex
1 O 12
2 O 10
3 O 24
4 O 29
5 O 30
6 O 18
7 O 32
8 O 26
9 F3 9
10 F3 9
11 F3 16
12 F3 4
13 S3 30
14 S3 7
15 S3 21
16 S3 9
17 F6 16
18 F6 10
19 F6 18
20 F6 18
21 S6 18
22 S6 24
23 S6 12
24 S6 19
25 F9 10
26 F9 4
27 F9 4
28 F9 5
29 S9 17
30 S9 7
31 S9 16
32 S9 17library(ggplot2)
# if (!require("ggplot2")) install.packages("ggplot2")
p13 <-
ggplot(data = df13, mapping = aes(x = Trt, y = ScabIndex)) +
geom_point() +
geom_boxplot() +
labs(x = "Treatment", y = "Scab Index") +
theme_bw()
print(p13)fm13 <- lm(formula = ScabIndex ~ Trt, data = df13)
summary(fm13)
Call:
lm(formula = ScabIndex ~ Trt, data = df13)
Residuals:
Min 1Q Median 3Q Max
-12.625 -4.844 0.625 3.594 13.250
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 22.625 2.369 9.549 8.08e-10 ***
TrtF3 -13.125 4.104 -3.198 0.003734 **
TrtS3 -5.875 4.104 -1.432 0.164666
TrtF6 -7.125 4.104 -1.736 0.094858 .
TrtS6 -4.375 4.104 -1.066 0.296601
TrtF9 -16.875 4.104 -4.112 0.000372 ***
TrtS9 -8.375 4.104 -2.041 0.051977 .
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 6.702 on 25 degrees of freedom
Multiple R-squared: 0.4641, Adjusted R-squared: 0.3355
F-statistic: 3.608 on 6 and 25 DF, p-value: 0.01026anova(fm13)Analysis of Variance Table
Response: ScabIndex
Df Sum Sq Mean Sq F value Pr(>F)
Trt 6 972.34 162.057 3.6081 0.01026 *
Residuals 25 1122.88 44.915
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Randomized Complete Block Design (RCBD)
- Used when experimental material is heterogenous in one direction
Example
Yield : Yield of barley, SoilType : Soil Type, and Trt : 5 sources and a control
Yield <- c(
32.1, 35.6, 41.9, 35.4
, 30.1, 31.5, 37.1, 30.8
, 25.4, 27.4, 33.8, 31.1
, 24.1, 33.0, 35.6, 31.4
, 26.1, 31.0, 33.8, 31.9
, 23.2, 24.8, 26.7, 26.7
)
SoilType <- gl(n = 4, k = 6, length = 4*6
, labels = c("I", "II", "III", "IV"), ordered = FALSE)
Trt <- gl(n = 6, k = 1, length = 4*6
, labels = c("(NH4)2SO4", "NH4NO3", "CO(NH2)2", "Ca(NO3)2", "NaNO3", "Control"), ordered = FALSE)
df14 <- data.frame(Yield, SoilType, Trt)
df14 Yield SoilType Trt
1 32.1 I (NH4)2SO4
2 35.6 I NH4NO3
3 41.9 I CO(NH2)2
4 35.4 I Ca(NO3)2
5 30.1 I NaNO3
6 31.5 I Control
7 37.1 II (NH4)2SO4
8 30.8 II NH4NO3
9 25.4 II CO(NH2)2
10 27.4 II Ca(NO3)2
11 33.8 II NaNO3
12 31.1 II Control
13 24.1 III (NH4)2SO4
14 33.0 III NH4NO3
15 35.6 III CO(NH2)2
16 31.4 III Ca(NO3)2
17 26.1 III NaNO3
18 31.0 III Control
19 33.8 IV (NH4)2SO4
20 31.9 IV NH4NO3
21 23.2 IV CO(NH2)2
22 24.8 IV Ca(NO3)2
23 26.7 IV NaNO3
24 26.7 IV Controllibrary(ggplot2)
# if (!require("ggplot2")) install.packages("ggplot2")
p14 <-
ggplot(data = df14, mapping = aes(x = Trt, y = Yield)) +
geom_point() +
geom_boxplot() +
labs(x = "Treatment", y = "Yield") +
theme_bw()
print(p14)fm14 <- lm(formula = Yield ~ SoilType + Trt, data = df14)
summary(fm14)
Call:
lm(formula = Yield ~ SoilType + Trt, data = df14)
Residuals:
Min 1Q Median 3Q Max
-7.0208 -2.4229 0.0792 2.1354 6.7958
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 35.354 2.826 12.511 2.44e-09 ***
SoilTypeII -3.500 2.664 -1.314 0.2087
SoilTypeIII -4.233 2.664 -1.589 0.1329
SoilTypeIV -6.583 2.664 -2.471 0.0259 *
TrtNH4NO3 1.050 3.263 0.322 0.7521
TrtCO(NH2)2 -0.250 3.263 -0.077 0.9399
TrtCa(NO3)2 -2.025 3.263 -0.621 0.5442
TrtNaNO3 -2.600 3.263 -0.797 0.4380
TrtControl -1.700 3.263 -0.521 0.6100
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 4.615 on 15 degrees of freedom
Multiple R-squared: 0.3512, Adjusted R-squared: 0.005215
F-statistic: 1.015 on 8 and 15 DF, p-value: 0.4653anova(fm14)Analysis of Variance Table
Response: Yield
Df Sum Sq Mean Sq F value Pr(>F)
SoilType 3 133.62 44.539 2.0915 0.1443
Trt 5 39.31 7.862 0.3692 0.8618
Residuals 15 319.43 21.295
Latin Square Design
- Used when experimental material is heterogenous in two perpendicular directions
Example
The following table shows the field layout and yield of a 5×5 Latin square experiment on the effect of spacing on yield of millet plants. Five levels of spacing were used. The data on yield (grams/plot) was recorded and is given below:
Yield <- c(
257, 230, 279, 287, 202,
245, 283, 245, 280, 260,
182, 252, 280, 246, 250,
203, 204, 227, 193, 259,
231, 271, 266, 334, 338
)
Row <- factor(x = rep(x = 1:5, each = 5))
Col <- factor(x = rep(x = 1:5, times = 5))
Trt <- c(
"B", "E", "A", "C", "D"
, "D", "A", "E", "B", "C"
, "E", "B", "C", "D", "A"
, "A", "C", "D", "E", "B"
, "C", "D", "B", "A", "E"
)
df15 <- data.frame(Yield, Row, Col, Trt)
df15 Yield Row Col Trt
1 257 1 1 B
2 230 1 2 E
3 279 1 3 A
4 287 1 4 C
5 202 1 5 D
6 245 2 1 D
7 283 2 2 A
8 245 2 3 E
9 280 2 4 B
10 260 2 5 C
11 182 3 1 E
12 252 3 2 B
13 280 3 3 C
14 246 3 4 D
15 250 3 5 A
16 203 4 1 A
17 204 4 2 C
18 227 4 3 D
19 193 4 4 E
20 259 4 5 B
21 231 5 1 C
22 271 5 2 D
23 266 5 3 B
24 334 5 4 A
25 338 5 5 Elibrary(ggplot2)
# if (!require("ggplot2")) install.packages("ggplot2")
p15 <-
ggplot(data = df15, mapping = aes(x = Trt, y = Yield)) +
geom_point() +
geom_boxplot() +
labs(x = "Treatment", y = "Yield") +
theme_bw()
print(p15)fm15 <- lm(formula = Yield ~ Row + Col + Trt, data = df15)
summary(fm15)
Call:
lm(formula = Yield ~ Row + Col + Trt, data = df15)
Residuals:
Min 1Q Median 3Q Max
-44.68 -12.48 1.12 16.52 54.92
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 240.08 23.43 10.247 2.75e-07 ***
Row2 11.60 20.55 0.565 0.5828
Row3 -9.00 20.55 -0.438 0.6692
Row4 -33.80 20.55 -1.645 0.1259
Row5 37.00 20.55 1.801 0.0969 .
Col2 24.40 20.55 1.187 0.2580
Col3 35.80 20.55 1.742 0.1070
Col4 44.40 20.55 2.161 0.0516 .
Col5 38.20 20.55 1.859 0.0877 .
TrtB -7.00 20.55 -0.341 0.7393
TrtC -17.40 20.55 -0.847 0.4137
TrtD -31.60 20.55 -1.538 0.1500
TrtE -32.20 20.55 -1.567 0.1431
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 32.49 on 12 degrees of freedom
Multiple R-squared: 0.6536, Adjusted R-squared: 0.3073
F-statistic: 1.887 on 12 and 12 DF, p-value: 0.1426anova(fm15)Analysis of Variance Table
Response: Yield
Df Sum Sq Mean Sq F value Pr(>F)
Row 4 13601.4 3400.3 3.2212 0.05164 .
Col 4 6146.2 1536.5 1.4556 0.27581
Trt 4 4156.6 1039.1 0.9844 0.45229
Residuals 12 12667.3 1055.6
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1